Postprocessing fMRI¶
When performing functional connectivity analysis, there are several additional processing steps that need to be taken after the minimal preprocessing of fMRIPrep. clpipe implements these steps in Python, and a fMRIprep preprocessed dataset can be postprocessed using the following command:
Usage: fmri_postprocess [OPTIONS] [SUBJECTS]...
This command runs an fMRIprep'ed dataset through additional processing, as
defined in the configuration file. To run specific subjects, specify their
IDs. If no IDs are specified, all subjects are ran.
Options:
-config_file PATH Use a given configuration file. If left blank, uses
the default config file, requiring definition of
BIDS, working and output directories.
-target_dir DIRECTORY Which fmriprep directory to process. If a
configuration file is provided with a BIDS
directory, this argument is not necessary. Note,
must point to the ``fmriprep`` directory, not its
parent directory.
-target_suffix TEXT Which file suffix to use. If a configuration file
is provided with a target suffix, this argument is
not necessary. Defaults to "preproc_bold.nii.gz"
-output_dir DIRECTORY Where to put the postprocessed data. If a
configuration file is provided with a output
directory, this argument is not necessary.
-output_suffix TEXT What suffix to append to the postprocessed files.
If a configuration file is provided with a output
suffix, this argument is not necessary.
-task TEXT Which task to postprocess. If left blank, defaults
to all tasks.
-TR TEXT The TR of the scans. If a cofig file is not
provided, this option is required. If a config file
is provided, this information is found from the
sidecar jsons.
-processing_stream TEXT Optional processing stream selector.
-log_dir DIRECTORY Where to put HPC output files. If not specified,
defaults to <outputDir>/batchOutput.
-beta_series Flag to activate beta-series correlation
correlation. ADVANCED METHOD, refer to the
documentation.
-submit Flag to submit commands to the HPC.
-batch / -single Submit to batch, or run in current session. Mainly
used internally.
-debug Print detailed processing information and traceback
for errors.
--help Show this message and exit.
Processing Checker¶
clpipe has a convenient function for determining which scans successfully made it through both preprocessing using fMRIprep and postprocessing.
Usage: fmri_process_check [OPTIONS]
This command checks a BIDS dataset, an fMRIprep'ed dataset and a
postprocessed dataset, and creates a CSV file that lists all scans across
all three datasets. Use to find which subjects/scans failed processing.
Options:
-config_file FILE The configuration file for the current data processing
setup. [required]
-output_file TEXT Path and name of the output CSV. Defaults to config
file directory and "Checker-Output.csv"
-debug Print traceback and detailed processing messages.
--help Show this message and exit.
This command will create a csv file listing all scans found in the BIDS dataset, and corresponding scans in the fMRIprep dataset and the postprocessed dataset.
For a description of the various postprocessing steps, along with references, please see the following documentation:
Nuisance Regression
Frequency Filtering
Scrubbing
Spectral Interpolation
fmri_postprocess2¶
New to clpipe v1.5, the command fmri_postprocess2 combines the functionality of fmri_postprocess and glm_setup into a unified postprocessing stream.
Usage: fmri_postprocess2 [OPTIONS] [SUBJECTS]...
Options:
-config_file FILE Use a given configuration file. [required]
-fmriprep_dir DIRECTORY Which fmriprep directory to process. If a
configuration file is provided with a BIDS
directory, this argument is not necessary. Note,
must point to the ``fmriprep`` directory, not its
parent directory.
-output_dir DIRECTORY Where to put the postprocessed data. If a
configuration file is provided with a output
directory, this argument is not necessary.
-processing_stream TEXT Specify a processing stream to use defined in your
configuration file.
-log_dir DIRECTORY Path to the logging directory.
-index_dir DIRECTORY Give the path to an existing pybids index database.
-refresh_index Refresh the pybids index database to reflect new
fmriprep artifacts.
-batch / -no-batch Flag to create batch jobs without prompt.
-submit Flag to submit commands to the HPC without prompt.
-debug Print detailed processing information and traceback
for errors.
--help Show this message and exit.
This command allows for flexible creation of processing streams. The order of processing steps and their specific implementations can be modified in the configuration file. Any temporally-relevant processing steps can also be applied to each image’s corresponding confounds file. fmri_postprocess2 caches its processing intermediaries in a working directory, which allows quick re-runs of pipelines with new parameters.
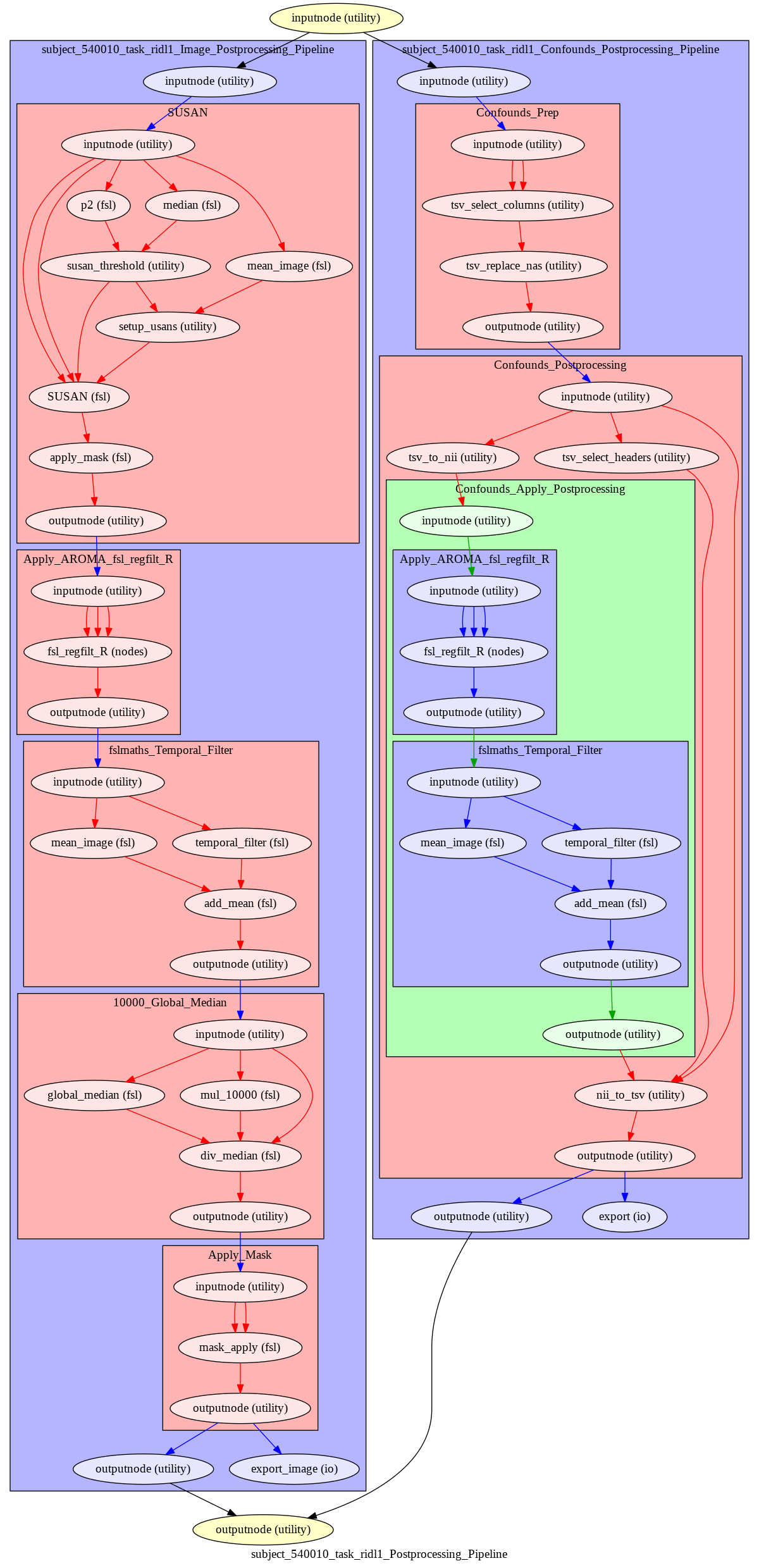
This command will also output a detailed processing graph for each processing stream.
Available processing steps:
Temporal Filtering
Intensity Normalization
Spatial Smoothing
AROMA Regression
Confound Regression
Apply Mask
Resample
Trim Timepoints
Configuration Setup¶
This command requires a new configuration block - if you using an existing clpipe project, you will have to insert this json into your configuration file. Otherwise, this block will be included when running “project setup.”
"PostProcessingOptions2": {
"WorkingDirectory": "",
"WriteProcessGraph": true,
"TargetImageSpace": "MNI152NLin2009cAsym",
"TargetTasks": [],
"TargetAcquisitions": [],
"ProcessingSteps": [
"SpatialSmoothing",
"TemporalFiltering",
"IntensityNormalization",
"ApplyMask"
],
"ProcessingStepOptions": {
"TemporalFiltering": {
"Implementation":"fslmaths",
"FilteringHighPass": 0.008,
"FilteringLowPass": -1,
"FilteringOrder": 2
},
"IntensityNormalization": {
"Implementation": "10000_GlobalMedian"
},
"SpatialSmoothing": {
"Implementation": "SUSAN",
"FWHM": 6
},
"AROMARegression":{
"Implementation": "fsl_regfilt_R"
},
"Resample":{
"ReferenceImage": "SET REFERENCE IMAGE"
},
"TrimTimepoints": {
"FromEnd": 0,
"FromBeginning": 0
},
"ConfoundRegression": {
"Implementation": "afni_3dTproject"
}
},
"ConfoundOptions": {
"Columns": [
"csf", "csf_derivative1", "white_matter", "white_matter_derivative1"
],
"MotionOutliers": {
"Include": true,
"ScrubVar": "framewise_displacement",
"Threshold": 0.9,
"ScrubAhead": 0,
"ScrubBehind": 0,
"ScrubContiguous": 0
}
},
"BatchOptions": {
"MemoryUsage": "20000",
"TimeUsage": "2:0:0",
"NThreads": "1"
}
}
PostProcessingOptions:Options for configuring post-fmriprep processing steps.WorkingDirectory:Directory for caching intermediary processing files.WriteProcessGraph:Set ‘true’ to write a processing graph alongside your output.TargetImageSpace:Which space to use from your fmriprep output. This is the value that follows “space-” in the image file names.TargetTasks:Which tasks to use from your fmriprep output. This is the value that follows “task-” in the image file names. Leave blank to target all tasks.TargetAcquisitions:Which acquisitions to use from your fmriprep output. This is the value that follows “acq-” in the image file names. Leave blank to target all acquisitions.ProcessingSteps:The default list of processing steps to use. Processing will follow the order of this list.ProcessingStepOptions:The default processing options for each step.TemporalFiltering:Apply temporal filtering to the image data. Also be applied to confounds.Implementation:Currently limited to “fslmaths”FilteringHighPass:High pass frequency for filtering. Defaults to .08 Hz. Set to -1 to remove high pass filtering.FilteringLowPass:Low pass frequency for filtering. Defaults to no filter (-1). Set to -1 to remove low pass filtering.FilteringOrder:Order of filter. Defaults to 2.
IntensityNormalization:Apply intensity normalization to the image data.Implementation:Currently limited to “10000_GlobalMedian”
SpatialSmoothing:Apply spatial smoothing to the image data.Implementation:Currently limited to “SUSAN”FWHM:The size of the smoothing kernel. Specifically the full width half max of the Gaussian kernel. Scaled in millimeters.
AROMARegression:Regress out AROMA artifacts from the image data. Also be applied to confounds.Implementation:Currently limited to “fsl_regfilt_R”
Resample:Resample the image into a new space.TrimTimepoints:Trim timepoints from the beginning or end of an image. Also be applied to confounds.FromEnd:Number of timepoints to trim from the end of each image.FromBeginning:Number of timepoints to trim from the beginning of each image.
ConfoundRegression:Regress out the confound file values from your image. If any other processing steps are relevant to the confounds, they will be applied first.Implementation:Currently limited to “afni_3dTproject”
ConfoundOptions:The default options to apply to the confounds files.Columns:A list containing a subset of confound file columns to use from each image’s confound file.MotionOutliers:Options specific to motion outliers.Include:Set ‘true’ to add motion outlier spike regressors to each confound file.ScrubVar:Which variable in the confounds file should be used to calculate motion outliers, defaults to framewise displacement.Threshold:Threshold at which to flag a timepoint as a motion outlier, defaults to .9ScrubAhead:How many time points ahead of a flagged time point should be flagged also, defaults to 0.ScrubBehind:If a timepoint is scrubbed, how many points before to remove. Defaults to 0.ScrubContiguous:How many good contiguous timepoints need to exist. Defaults to 0.
BatchOptions:The batch settings for postprocessing.MemoryUsage:How much memory to allocate per job.TimeUsage:How much time to allocate per job.NThreads:How many threads to allocate per job.
Processing Streams Setup¶
By default, the output from running fmri_postprocess2 will appear in your clpipe folder at data_postproc2/smooth_filter_normalize, reflecting the defaults from PostProcessingOptions2.
However, you can utilize the power of processing streams to deploy multiple postprocessing streams. Each processing stream you define your config file’s ProcessingStreams block will create a new output folder named after the ProcessingStream setting.
Within each processing stream, you can override any of the settings in the main PostProcessingOptions2 section. For example, in the follow json snippet, the first processing stream will only pick “rest” tasks and defines its own set of processing steps. The second stream does the same thing, but specifies a filtering high pass by overriding the default value of -1 with .009.
...
"ProcessingStreams": [
...
{
"ProcessingStream": "smooth_aroma-regress_filter-butterworth_normalize",
"PostProcessingOptions": {
"TargetTasks": [
"rest"
],
"ProcessingSteps": [
"SpatialSmoothing",
"AROMARegression",
"TemporalFiltering",
"IntensityNormalization",
"ApplyMask"
]
}
},
{
"ProcessingStream": "smooth_aroma-regress_filter-high-only_normalize",
"PostProcessingOptions": {
"TargetTasks": [
"rest"
],
"ProcessingSteps": [
"SpatialSmoothing",
"AROMARegression",
"TemporalFiltering",
"IntensityNormalization",
"ApplyMask"
],
"ProcessingStepOptions": {
"TemporalFiltering": {
"FilteringHighPass": .009
}
}
}
},
...
To run a specific stream, give the -processing_stream stream option of fmri_postprocess2 the name of the stream:
fmri_postprocess2 -config_file clpipe_config.json -processing_stream smooth_aroma-regress_filter-butterworth_normalize -submit